Breeding dogs with a purpose- and actually achieving the goal of dogs capable of such a purpose- has always been a blend of art and science. In recent years, the science part of that equation has grown and developed in ways we may have never even imagined possible.
When we think of the science of breeding, a lot of it is centered around canine health, soundness and longevity- and rightly so, because no matter what the purpose is of a particular breed or specific breeding program, the one thing every good breeder wants is healthy offspring. Some measures of health are based on more definitive science than others- for example, for some heritable diseases, researches have pinpointed the specific mutated gene that causes the disease. Using this information, breeders can run very simple and affordable tests on their breeding dogs in order to avoid producing puppies affected by that disease. Brilliant!
Even more recently, breeders have teamed up with researchers from the University of California Davis to figure out how we can determine the relatedness between two dogs as well as the genetic value of a dog in comparison to other members of the same breed. In the past, we’ve had to rely on pedigrees (looking for common ancestors) or using pedigree software to calculate the Coefficient of Inbreeding- which is an estimate of how related two dogs may be to each other using a mathematical formula. Both of these methods are limited because as we all know, puppies from the same litter may not inherit exactly the same set of genes from each parent dog, and so within a few generations, some particular genes from a dam or sire may be very prolific in their descendants, or gone entirely, depending on which puppies in a litter are bred! There is no way to know which are inherited in a specific dog for genes that don’t have a specific test or show outward clues, but it underlines that pedigree research and calculations of COI are very limited at best in determining the relationship between two dogs on a genetic level.
So- why does this matter anyway? Breeders work very hard to breed dogs that are consistently healthy, with good temperaments, and well suited for their ‘job’, whatever that may be for the breed. We plan breedings in an attempt to maintain the strengths of the individual parent dogs, and improve upon their weaknesses. But consistency and improvement are opposing forces…. you can’t improve something without variation, and you don’t get consistency if there is too much variation. Additionally, in order to breed ‘the best of the best’, potentially valuable genes are lost from dogs who could otherwise contribute positive features to the breed. Think about a dog who may have a tail that is too long for it’s breed so is overlooked as a breeding prospect in the effort to only use the very best dogs for breeding but…. what if that dog was the only one of it’s littermates to inherit a genetic combination that was resistant to cancer? What if he lived to be 18 years old while all of his brothers died at 12? Some desirable features are very difficult to select for or aren’t known until it’s too late, and with purebred dogs, we only get fewer genes with each generation. New blood is never added; so ensuring we don’t throw out dogs who may be valuable genetic contributors is super important.
Some breeders use line-breeding techniques to try to gain consistency. After much research, the breeder identifies a particularly strong individual or line of dogs, and breeds back to those relations in hopes of increasing the proportion of puppies in each generation that have the desirable traits of this spectacular ancestor. Over time, variability decreases. However, one is never truly able to select for just one trait at a time- a dog’s genome codes for thousands of different traits- and it is possible that as a side effect of seeking out one or a few valued traits in a family, that we may accidentally also increase the undesirable traits that were hidden (recessive) in the original dog, making those also more frequent in later generations and hard to avoid due to the lack of variation that remains.
Some breeders use out-crossing techniques to try to make improvements. Choosing an unrelated dog who is particularly strong in a trait or set of characteristics where the original breeding dogs are not as strong, and hoping that more of the puppies will get this desirable trait than not. However when one breeds dogs who are distinctively different, you are introducing more variability and it is possible that some puppies get the best traits of both parents, while others get the worst of them. Typically, from a health standpoint, variability is presumed to be helpful in general, so long as the variation that is introduced isn’t hiding some dark dirty health secrets. In purebred breeding programs, some breeds have such a small number of actively breeding dogs that it is very difficult to identify which dogs may be less related, based on pedigree research alone.
Some breeders use a blend of line-breeding and outcrossing in attempt to balance the risks and benefits of each. Ah! It gets tricky, particularly within the confines of a pure breed, and when your options for breeding candidates for the next generation rely at least in part on the decisions made by other breeders within the breed! I may want to breed dogs who are unrelated to Popular Sire Z, but if everyone of my peers breeds to this dog next year, it’s going to be super tough going forward to find dogs who aren’t descendants of Z! By default, some breeders are going to end up line-breeding on this dog whether it is a good decision or not, due to the lack of options presented by this mass-breeding genetic bottleneck.
And this is where the new science really has the ability to pay off. BetterBred is a company that has teamed up with UC Davis to help breeders understand the applications of genetic profiling and it’s usefulness for planning litters. Most of us are already well aware of how to use specific gene tests to avoid disease mutations, such as Exercise Induced Collapse. But this is new and a little more complicated!
The testing through Betterbred scans the dog’s genome and records the alleles found at dozens of specific loci. These are then compared to the other breed members in the database. There are different measuring parameters which can identify a dog who has genes who are very common to the other dogs in the database (ie related to most of them), dogs who have very little variation in their own alleles (ie are quite inbred), and dogs who have genes who may be less common within the breed, and worthy of preserving for the genetic health going forward.
What does this mean exactly for breeders? This means that breeders who want to use line-breeding can identify if their dogs actually are related to each other (ie have inherited similar genes from the common ancestor they are wanting to duplicate). It also allows breeders who are wanting to outcross to use dogs who actually do have different genetic makeup- without necessarily having to find an entirely divers pedigree. And it allows breeders who want to ensure that there is an option to preserve unique, potentially important uncommon genes, by using dogs in their breeding program who have a less common genetic makeup.
For us here at Eromit, we really want to produce dogs who have the consistent ability to be strong working and performance dogs but we don’t want to sacrifice health to do it. In the past, we have bred ‘like to like’ in terms of choosing dogs with similar temperaments, drive, and structure, but from as unrelated pedigrees as we could find within the working Lab worlds. (Yes, there is a little more to it than this in terms of specific selection traits, but this is oversimplified gist). NOW, however, the use of Betterbred can really help confirm whether we are achieving what we have set out to do. Instead of having to look for dogs who have wildly different pedigrees in order to avoid inbreeding, we can actually test the dogs we are interested in breeding to each other, to confirm whether or not they show a close relationship or not.
Here is an example. When we chose Harvey as a potential stud dog, one of the factors we were considering was that we wanted a dog who was not closely related to our current dogs. We want to ensure that we are not losing variation that helps prevent unintended accumulation of recessive disease alleles, but it is very hard when you already have a line that contains very popular breeding dogs, especially with chocolate labs. Most of the really talented chocolate dogs in the field world are descendants of the same few dogs. Luckily for us, we were able to find Harvey whose pedigree is free from nearly all of those popular chocolate sires while still maintaining generations of health tested, working verified well-bred dogs.
Using the BetterBred program, we can simulate a breeding of Jet to Harvey. I chose Jet for this example because her pedigree is a little less common than some of our other girls and incorporates a line that was not widely bred. Jet is a small, soundly built female with wonderful work ethic, a great temperament, and that coveted off-switch, so keeping dogs like her in the gene pool seems like a very good decision, particularly given her less common ancestry. It is easy to find unrelated dogs to breed to Jet, but that ease could be lost in the next generation by breeding her to a very common dog or line.

So according to the actual genetic makeup of Jet and Harvey, and not some hypothetical estimate of inheritance- they are as unrelated as two labs can get. Excellent! Compared to the other Labs in the database, the litter would be expected to produce puppies who are quite different in genetic makeup (which means more potential unrelated breeding partners for them- that is also excellent). For each individual puppy, the range of expected inbreeding – or internal relatedness- is also very low, with most of the puppies expected to be much less ‘inbred’ than the average lab. That is an excellent target for purebred dogs, as it means we are utilizing the full scope of the genes remaining in the breed.
Compare this breeding of Jet to Buzz. When we chose Buzz, we were equally looking for a dog that was unrelated based on pedigree to our current girls, but without going outside of the field trial type of Labs. This is much more doable in black and yellow dogs than in chocolate; though there is quite a bit of ‘popular sire syndrome’ dogs in these colors too, there are just that many more well bred yellow and black dogs to choose from and it is still currently possible to avoid pedigrees with specific popular dogs.
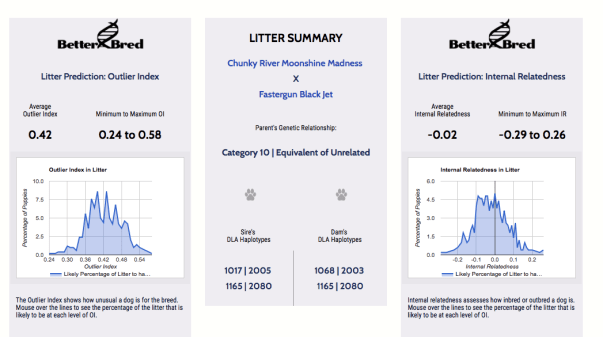
Interestingly, even though Buzz is all field lines, and Harvey is not, both sires are entirely unrelated to Jet. Crossing Jet to Buzz produces similar levels of inbreeding (or more accurately, lack of inbreeding) and quite similar levels of unrelatedness to other dogs in the database. Using the BetterBred tool, we can get predictions of the levels of inbreeding in litters of both combinations, and use that in addition to our usual selection tools to make an informed decision about which sire to choose as a best match.
In this case, the BetterBred tool did not really act as a tie-breaker but did confirm that we would not end up with highly inbred puppies from either sire option. We ended up choosing Buzz as a sire for Jet’s last litter, as it provided the most ‘like to like’ in terms of temperament and drive, with complementary structure & health testing, while maintaining genetic diversity for the goal of health and longevity for the next generation.
Here is a second example.
Avi is one of our females that is lightly line bred on on our foundation female “Nestle” (very lightly- Nestle is Avi’s Grandma on her mother’s side, and her great-great-great gramma on her sire’s side). Avi also has several popular field trial line sire woven in there which means that she is related to a lot of different lab lines.
Avi has been bred twice to Harvey. Here is what their BetterBred analysis looks like: 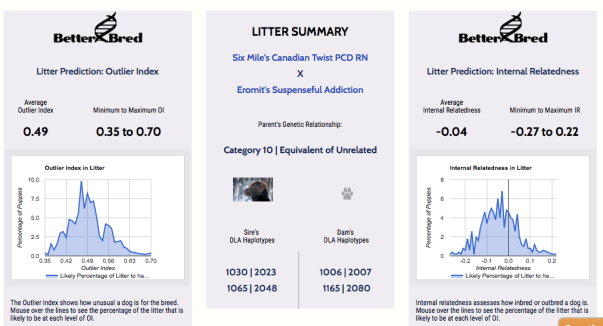
As expected based on pedigree, Avi and Harvey are genetically unrelated. The puppies they produced were not highly related to most of the other dogs in the database and their range of inbreeding averages lower than the breed average, though there is quite a bit of variation possible there. The variation did show up in the puppies in other ways too, as some of them very much took after their sire’s side, and some took very much after the mother’s side (in terms of looks, temperament, or both) and some were the expected ‘blend’.
The science here has potential use for this next generation- if I was so inclined, I could have tested the puppies and kept the puppy that was on the lower end of the internal relatedness scale in order to give me the most options for maintaining variation in the next generation (or used that measurement as a tie-breaker between two puppies that I otherwise liked), or I could have chosen one with a higher degree of inbreeding -which would have meant more potential suitor options in the next generation.
For Avi’s next litter, we are considering a couple of other sires in the hopes of choosing one that is a little more similar to Avi without risking a tight inbreeding. Remember this is going to be trickier because of how many popular lines are contained within her pedigree.
How about Buzz?

Well- how about that. Despite only one common popular ancestor in their 5 generation pedigrees, Avi and Buzz are reasonably related on the genetic front. According to BetterBred, this is comparable to a ‘half first cousins’ breeding’. Looking at the numbers though, the differences between this litter, and the one above to Harvey, are not extreme. The range of internal relatedness is remarkably similar. The puppies from this litter are more likely to be related to the average lab than puppies sired by Harvey.
Here is one more potential sire:

“Coulee” is very similar in many ways to Buzz, though an entirely different pedigree. This is reflected in the fact that Avi and Coulee are genetically unrelated. The puppies from his breeding would be even less common than Harvey’s litter, and have a similar range of internal relatedness to Buzz’s litter (actually, even little less variation).
What does this really mean for our breeding program? Well, there are a few options here (considering only these three sires for illustration purposes). We really liked our Avi x Harvey puppies but there was more variation there than we normally see. In order to gain a bit of consistency in the next litter, we could choose Coulee as a sire instead. Genetic analysis of the puppies in the litter could help us decide to keep either a puppy with a higher internal relatedness score (perhaps a future breeding candidate to go back to Harvey?) or a lower IR score (to breed back to Buzz or similar)?
We are just scratching the surface in understanding the usefulness of this type of science to aid in breeding decisions, and the potential will grow as more Labradors are tested and enter the database for comparison. It is possible that this sort of testing will eventually be one of the primary tools used by purebred breeders to ensure we breed dogs with the traits we desire, without losing genetic diversity important to the long term health of the breed OR the individual dogs.
Hopefully, I’ve not oversimplified this too much. If you are a breeder, or otherwise interested in the details of the specific measurements in these charts, I would encourage you to check out the free online course offered by BetterBred which helps explain things with much more precision. https://www.betterbred.com/product/introduction-to-genetics-with-betterbred/
Recent Comments